
— Benefits
15x acceleration, 6-8x less energy consumption, 10-15x TCO
Data-intensive application developers are looking for new solutions to gain an order of magnitude in efficiency at server-level. The legacy processing-centric model cannot help, even with the multiplication of cores because the limiting factor is the data transfer bottleneck between the memory and the computing units.
UPMEM PIM DRAM solution is a new class of memory-centric solutions that utilizes thousands of parallel processors. It has already demonstrated over 15x performance improvement compared to a standard server and for multiple use cases.
UPMEM PIM DRAM solution is based on UPMEM DPU processor instances integrated into the DRAM memory chips, where the data is located. UPMEM DPU processor is both general purpose and optimized for data computing. The PIM chips (combining DRAM and UPMEM DPUs) are assembled on DIMM modules, and plugged into the memory slots of the server.
PIM in DRAM is very competitive because its production cost is mostly sitting on DRAM processes one, which is half the price of a logic one.
For a full review of the publicly released work on UPMEM PIM, check our available resources.
Performance x15

— Genomics
Running genomics algorithms in minutes instead of hours
The main genomics operations are extremely data-intensive applications. A single human genome sequencing produces ~190GB of data to process.
Mapping and Variant calling of DNA chain fragments against a reference genome can be accelerated over 100x against BWA-GATK reference or about identical to other accelerated pipelines, reducing standard processing times from days or hours to a few minutes.
Combining those genomics operations results in virtually real-time personalised medicine. It also reduces TCO by up to a factor 12 when compared to other acceleration solutions and at identical throughput thus making recent progress in genomics accessible to all.
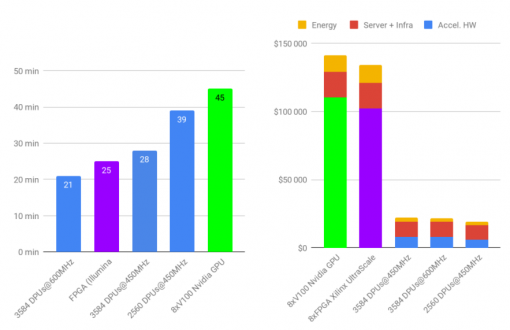
The INRIA’s paper for the BIBM2020 conference about Mapping & Variant Calling, showing similar time to current accelerator (GPU and FPGA) but with x8-12 TCO gain and x6-8 energy reduction, can be found here
Mapping and Variant calling of human genomes x 12 TCO
BLAST protein chain queries x25
Long read alignment (KSW2 implementation) x 15
— Database
Divide by 100 database index search response time, multiply throughout by 17
Searching strings of words in indexed document database is a massive and well identified application. Implementing such a workload on UPMEM PIM solution drastically leverages the thousands of DPU cores that can work in parallel for each request.
The work called UPIS demonstrates the ability of a PIM server to process over 60x more requests per second depending on the PIM configuration and with 2 orders of magnitude better latency than the equivalent x86 server equipped with DRAM. TCO and energy gains range between 40 times better than an x86 server.
Speed-up, energy gains and TCO gains of processing search requests via PIM UPIS app running on PIM equipped servers vs. via Lucene Apache running on the same server equipped with conventional DRAM memory, on the same database. Performance is considerably better when batching requests, while latency increases.
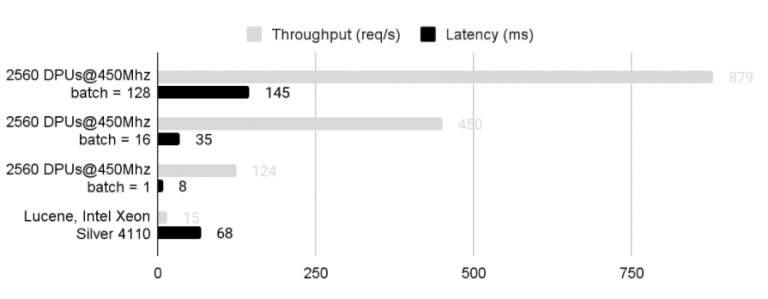
Response time x100 faster
Better throughput at marginal additional cost x8

— Many more use cases
Applications on UPMEM PIM are limitless
UPMEM cooperates with dozens of renowned labs and R&D centers around the world to constantly explore and benchmark new applications of PIM.
The list of use case is constantly expanding but advanced works have already identified great acceleration potentials for PIM in the following topics:
- Compression/Decompression (Snappy)
- Data analytics
- Machine Learning & AI (DLRM, Kmeans, CNN, Decision Trees, etc.)
- Graph Algorithms
- 3D image reconstruction & FFT
- n-step FM index
- Skyline (multi-feature preference query)
- Seismic reconstruction
Reach to us to know more about the ongoing work or benchmarks available or if you have a use case of your own that you would like to explore together.
— Start exploring PIM implementations
Check out the latest release from the PIM community
On UPMEM’s github, you will find numerous small applications and benchmarks:
The ETH Zurich under the guidance of Onur Mutlu, has released an extensive analysis of the UPMEM PIM architecture in their paper: “Benchmarking a New Paradigm: An Experimental Analysis of a Real Processing-in-Memory Architecture”
More than 16 core algorithms are benched on PIM and versus GPU and CPU: https://github.com/CMU-SAFARI/prim-benchmarks
The University of British Columbia has made available several of their algorithms in data analysis: from compression to Hyper Dimensional Computing:
